Making DrugBank Reproducible
If you’re reading my blog, there’s a pretty high chance you’ve used DrugBank, a database of drug-target interactions, drug-drug interactions, and other high-granularity information about clinically-studied chemicals. DrugBank has two major problems, though: its data are password-protected, and its license does not allow redistribution. Time to solve these problems once and for all.
I’d guess that most of the thousands of people who use DrugBank have run into the same issue as me to get the data: you have to navigate through the DrugBank site, create an account, log in, then download the data manually. There are some hints on the site on how this can be done through the shell, but I think it’s the unfortunate case that many people using bioinformatics resources just aren’t comfortable using the shell. Further, the file is zipped, which means that unzipping it requires further knowledge of the arcane arts of which shell programs and which flags to use with them.
Even after getting past this, users need to write their own programs that read the XML content of the file, extract the relevant parts, and put it somewhere for later. If you’re like me, you probably want to have as little to do with data in the XML format as possible, and you probably deleted the original zip archive along with the XML file after you got the data out of it that you actually wanted.
The trouble with the scenario described in the last two paragraphs is that any analysis done on the resulting files required lots of manual steps in the middle. These steps can’t be automated, and therefore the downstream analysis can’t be automated either. It’s highly likely for a research to have the time or motivation to do these steps exactly the same way as described in a paper, if it’s even described at all.
Because of DrugBank’s licensing rules, you can’t redistribute the data in the format that is necessary for downstream use either.
This is no small problem - hundreds of papers cite DrugBank per year. The hype of drug repositioning for COVID-19 no doubt helped in 2020, with 147 citations as of the time of writing this post on December 14th, 2020.
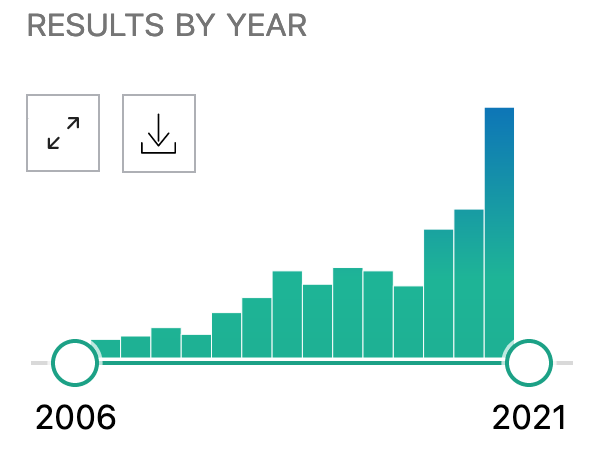
Search conducted on December 14th, 2020 with https://pubmed.ncbi.nlm.nih.gov/?term=drugbank.
It’s difficult to tell which, if any, of these efforts are meaningful. Without the ability to reproduce the steps taken to do analyses based on DrugBank, we can not even begin to evaluate these papers. Even further - imagine the intense pressure on the few famous scientists in network-based drug repositioning who receive the most requests to review more papers. How could we possibly expect these scientists to provide thorough reviews if it’s not easy (or possible, for that matter) for them to re-produce the analyses?
The Solution
The first time I put to writing the need to automate the acquisition of data was in my (still not accepted through peer review) article on the reproducible acquisition and conversion a wide range of biological databases into Biological Expression Language. In this article, we presented a suite of 50+ scripts which each downloaded, converted, and exported a biological database like DrugBank, GWAS Catalog, HIPPIE, and others into a common schema for networks biology downstream applications like signal transduction pathway analysis, knowledge graph embedding, and other methods. Of all of them, DrugBank remained the most vexing.
It’s time for a solution - for code that can help get us out of this rut. This code needs to be able to download the data from DrugBank, open it up, and help transform it all without asking the user to intervene too much.
Enter drugbank_downloader.
This is a Python library with three functions exactly for this. First, you have
to install it with:
$ pip install drugbank-downloader
Download A Specific Version
If you want your analysis to be reproducible, you should specify what version of
the database you want to use. Luckily, DrugBank does keep an
archive of old versions. The following code
describes how to use the drugbank_downloader.download_drugbank() function,
which asks for a version, a DrugBank username, and the corresponding DrugBank
password. You’re free to get the username and password into the script however
you want - usually loading from the environment using os.getenv() is a good
choice. Hard-coding them is not so much, because it could potentially violate
the terms and conditions of use of DrugBank, and just isn’t good practice in
general, especially if you reuse passwords. Get
LastPass.
import os
from drugbank_downloader import download_drugbank
username = ... # suggestion: load from environment with os.getenv('DRUGBANK_USERNAME')
password = ...
path = download_drugbank(version='5.1.7', username=username, password=password)
# This is where it gets downloaded: ~/.data/drugbank/5.1.7/full database.xml.zip
expected_path = os.path.join(os.path.expanduser('~'), '.data', 'drugbank', '5.1.7', 'full database.xml.zip')
assert expected_path == path.as_posix()
This script takes care of downloading the data and storing it at
~/.data/drugbank/5.1.7/full database.xml.zip. After it’s been downloaded once,
the function is smart and doesn’t need to download again. This is all taken care
of by the pystow behind the scenes. If you
specify an alternate version, it’s pretty obvious where in the filepath it
changes.
Download the Latest Version
If you want to automate running your code on the most recent version of
DrugBank, you can use the same function, but you’ll first have to install
bioversions with
pip install bioversions. This package’s job is to look up the most recent
version of varions biological databases. I’ll have a whole post on that later.
Then, you can modify the previous code slightly by simply removing the version
argument.
import os
from drugbank_downloader import download_drugbank
username = ... # suggestion: load from environment with os.getenv('DRUGBANK_USERNAME')
password = ...
path = download_drugbank(username=username, password=password)
# This is where it gets downloaded: ~/.data/drugbank/5.1.7/full database.xml.zip based on the latest
# version, as of December 14th, 2020.
expected_path = os.path.join(os.path.expanduser('~'), '.data', 'drugbank', '5.1.7', 'full database.xml.zip')
assert expected_path == path.as_posix()
Don’t Bother Unpacking - read full database.xml.zip Directly
Like I said before, it’s not enough just to get the file. It needs to be unzipped and opened. If you’re not familiar with doing this in python, you can use the following code:
import zipfile
from drugbank_downloader import download_drugbank
path = download_drugbank(username=..., password=...)
with zipfile.ZipFile(path) as zip_file:
with zip_file.open('full database.xml') as file:
pass # do something with the file
You don’t have time to remember this. Just use
drugbank_downloader.open_drugbank() instead:
from drugbank_downloader import open_drugbank
with open_drugbank(username=..., password=...) as file:
pass # do something with the file, same as above
There’s some magic using contextlib to make this work with the with
statement. Just keep reading, though…
Reading Drugbank’s XML
After you’ve opened the file, you probably want to read it with an XML parser like:
from xml.etree import ElementTree
from drugbank_downloader import open_drugbank
with open_drugbank(username=..., password=...) as file:
tree = ElementTree.parse(file)
root = tree.getroot()
You don’t have time to remember this either. Just use
drugbank_downloader.parse_drugbank() instead:
from drugbank_downloader import parse_drugbank
root = parse_drugbank(username=..., password=...)
DrugBank: Endgame
Once you’ve got the XML python object, you can write all of the code to
extract the parts you want, load them in a pandas.DataFrame, or do whatever
you’d like. If these steps are slow, you could even write code that caches it in
the middle. Below is my template for an expensive processing step.
from drugbank_downloader import parse_drugbank
def process_data():
...
def save_my_parsing_results(processed_data, file):
...
def load_my_parsing_results(path):
...
def get_processed_data():
cache_path = ...
if os.path.exists(cache_path):
processed_data = load_my_parsing_results(cache_path)
else:
root = parse_drugbank(username=..., password=...)
processed_data = process_data(root)
with open(cache_path, 'w') as file:
save_my_parsing_results(root, file)
return processed_data
I understand that most scientists aren’t trained as software engineers, nor are they incentivized to write code that’s reproducible, but I at least hope this code helps cut a few non-reproducible steps out of your science. Happy drug hunting!