Pathway Relationships
Domingo-Fernandez et al. published ComPath: An ecosystem for exploring, analyzing, and curating mappings across pathway databases. in 2018 describing the overlap between human pathways in KEGG, Reactome, and WikiPathways. A lot of the underlying machinery I developed to support this project has been improved since, and it’s time to spread the search to other organisms besides humans and other databases. This blog post is about some additional relation types needed to capture the relations between pathways appearing in these databases.
Like many of my blog posts, this one was inspired by a tweet. After the following discussion, I thought it would be good to better organize the ideas and elaborate.
#KEGG provides a namespace of non-species-specific pathway terms that all of its species-specific pathways can map to. @reactome could you provide the same?
— Charles Tapley Hoyt (@cthoyt) December 30, 2020
This blog post will follow apoptosis, one of the most ubiquitous pathways in biology that covers all manners of programed cell death. This blog post isn’t about the nitty-gritty difference between pathways, biological processes, and mechanisms - so we will consider all variants of apoptosis and apoptotic process effectively the same.
| Resource | Prefix | Identifier |
|---|---|---|
| Gene Ontology (GO) | go | GO:0006915 |
| Medical Subject Headings (MeSH) | mesh | D017209 |
| Kyoto Encyclopedia of Genes and Genomes (KEGG) | kegg.pathway | map04210 |
| NCI Thesaurus (NCIT) | ncit | C17557 |
KEGG, Reactome, and WikiPathways all provide human-specific variants of these pathways (below) as well as many other species, including both model organisms and not.
| Resource | Prefix | Identifier |
|---|---|---|
| KEGG | kegg.pathway | hsa04210 |
| Reactome | reactome | R-HSA-109581 |
| WikiPathways | wikipathways | WP254 |
Pathways are Equivalent
Two pathways are equivalent and can be represented with skos:exactMatch if
they both have the same species specificity. The following relationships are
between the non-species specific pathways for apoptosis:
| Subject | Predicate | Object |
|---|---|---|
| GO:0006915 | skos:exactMatch | mesh:D017209 |
| GO:0006915 | skos:exactMatch | kegg.pathway:map04210 |
| mesh:D017209 | skos:exactMatch | kegg.pathway:map04210 |
| … | … | … |
The following relationships are between the human-specific pathways for apoptosis in KEGG, Reactome, and WikiPathways:
| Subject | Predicate | Object |
|---|---|---|
| kegg.pathway:hsa04210 | skos:exactMatch | reactome:R-HSA-109581 |
| kegg.pathway:hsa04210 | skos:exactMatch | wikipathways:WP254 |
| wikipathways:WP254 | skos:exactMatch | reactome:R-HSA-109581 |
Similarly, the relationships between cow-specific (Bos Taurus; BTA) pathways for apoptosis in KEGG, Reactome, and WikiPathways:
| Subject | Predicate | Object |
|---|---|---|
| kegg.pathway:bta04210 | skos:exactMatch | reactome:R-BTA-109581 |
| kegg.pathway:bta04210 | skos:exactMatch | wikipathways:WP1018 |
| wikipathways:WP1018 | skos:exactMatch | reactome:R-BTA-109581 |
While equivalences begins to tame the ontology of pathways, it is missing links between the GO, MeSH, and NCIT terms to Reactome and WikiPathways.
Species-Specific Variant of a Pathway
GO, MeSH, NCIT, and many other nomenclatures do not contain species-specific
variants of their pathways. However, KEGG contains both a parent pathway,
prefixed with map and species-specific pathway, prefixed with their internal 3
or 4-letter species code.
| Subject | Predicate | Object |
|---|---|---|
| kegg.pathway:hsa04210 | speciesSpecific | kegg.pathway:map04210 |
| kegg.pathway:bta04210 | speciesSpecific | kegg.pathway:map04210 |
| … | … | … |
It should generally hold that when X speciesSpecific Y and
Y skos:exactMatch Z are true, X speciesSpecific Z. This allows KEGG to serve
as a bridge between the species-specific and non-species-specific pathway
worlds. However, Domingo-Fernandez et al. showed that there are huge
discrepancies between KEGG, Reactome, and WikiPathways, so there is still need
to curate/infer the same kinds relationships in Reactome and WikiPathways.
Unfortunately, Reactome and WikiPathways do not (yet) have parent terms for non-species-specific pathways. Asking about this was the point of the tweet that inspired this blog post. Because Reactome uses a standardized nomenclature where all variants of each pathway across species have the same numerical part to their identifier (e.g., R-HSA-109581 and R-BTA-109581), they could institute a similar parent nomenclature like KEGG’s. WikiPathways identifiers do not have this sort of regularity, but they have the benefit of being highly receptive to external input and improvements.
Side bar: I’ve seen an elegant solution for this in OBO that defines child terms with an intersection of the Relation Ontology relation RO:0002160 (only in taxon) to a given species and the parent term, but this is an unnecessarily complicated alternative for the goal of representing the relation between two entities.
Pathways are Orthologs
Two genes with similar evolutionary history and function appearing in two organisms are called orthologs. Orthology is incredibly important for studying biology because it allows us to make inferences about how human biology works by studying model organisms like mice and rats. There are several databases collecting orthology relationships, such as HomoloGene.
It follows that orthology could be applied to pathways as well. In fact, Reactome’s web interface already has a box below each pathway linking to the orthologous pathways as seen on https://reactome.org/content/detail/R-HSA-109581:
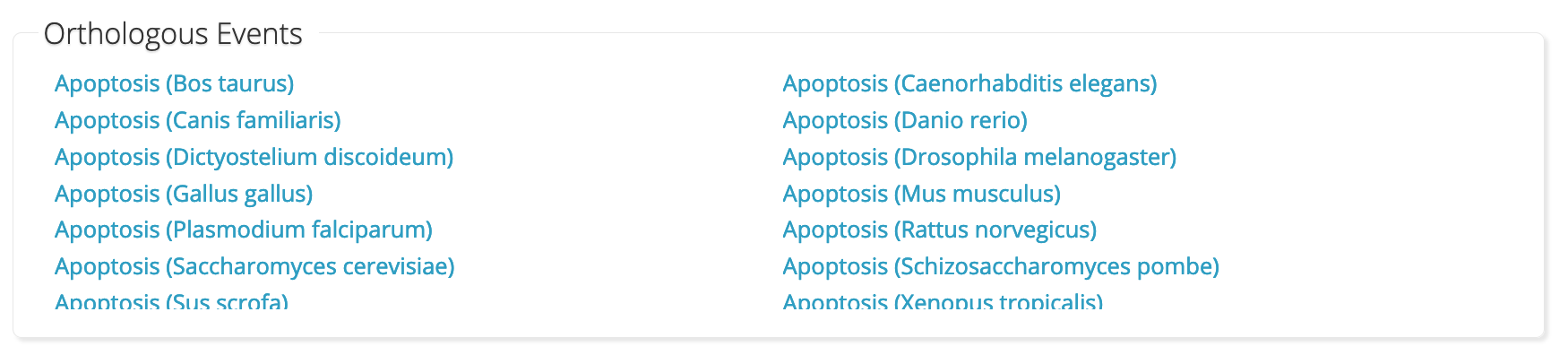
However, this information is not programatically available (AFAIK), and it is
not available for other databases like WikiPathways and KEGG. Therefore, we can
introduce a relationship orthology to start curating triples like:
| Subject | Predicate | Object |
|---|---|---|
| kegg.pathway:hsa04210 | orthology | kegg.pathway:bta04210 |
Orthology relationships effectively convey the same information as
speciesSpecific with the advantage that they do not require the addition of a
parent term. However, between N orthologous pathways, there will be a complete
subggraph of (1/2) _ N _ (N-1) edges (also called a clique in graph theory).
Depending on the downstream use case, these kinds of subgraphs can be
problematic.
Because kegg.pathway:hsa04210 skos:exactMatch reactome:R-HSA-109581, we can
infer reactome:R-HSA-109581 orthology kegg.pathway:bta04210. However, I think
it would be best to only curate orthology relationships within a given database
because it will increase the size (N) of the clique.
Pathway is About a Concept
KEGG, Reactome, and WikiPathways not only include pathways, but also other “maps” about specific topics such as diseases, families of proteins, and other biological entities.
For example, KEGG has an entry
kegg.pathway:hsa05010 entitled
Alzheimer disease - Homo sapiens (human). When using
Gilda to generate lexical matchings, the
MeSH entry mesh:D000544 (Alzheimer
Disease) appeared highly ranked. However, KEGG’s notion of pathway and MeSH’s
notion of a disease are not the same, and these two terms should not be
considered equivalent. For this case, not only KEGG but also Reactome and
WikiPathways, we can introduce a new relationship pathwayAbout. It turns out
that WikiPathways also has an Alzheimer’s disease “pathway” as well.
| Subject | Predicate | Object |
|---|---|---|
| kegg.pathway:hsa05010 | pathwayAbout | mesh:D000544 |
| wikipathways:WP2059 | pathwayAbout | mesh:D000544 |
Note that KEGG and WikiPathways both have specificity in their pathways for
organisms, but diseases in MeSH and other nomenclatures aren’t typically
stratified by their target organisms. Therefore, the mouse-specific Alzheimer’s
disease pathway in WikiPathways (wikipathways:WP2075) could also have the same
relationship.
Another example is opsins - a family of light-sensitive proteins. Reactome has a pathway reactome:R-HSA-419771 (Opsins) that is not the same as the MeSH entry mesh:D055355 (Opsins) describing the protein family.
There is specific interest in connecting disease maps appearing in pathway databases to the diseases themselves. WikiPathways has already begun doing this as can be seen on https://www.wikipathways.org/index.php/Pathway:WP2059.

It might be justified to propose an alternate relationship with more specific semantics. More information on various disease-specific curation projects outside major pathway databases can be found at https://disease-maps.org.
Disease-specific Variant of a Pathway
This is a bit of an afterthought, but it might be mentioning that there are places, like NeuroMMSig, that curate disease-specific variants of pathways. These would need their own dedicated relationships to connect to the “canonical” pathway and to the disease that they describe.
I have to give a huge shout-out to Daniel Domingo-Fernández, Josep Marín-Llaó, Carlos Bobis-Álvarez, and Yojana Gadiya who have done the curation in the ComPath project as well as Ben Gyori who laid the groundwork for improving the lexical mappings with the Gilda software as well as contributed tons of curations for MeSH-GO mappings.
There are still many disjoint resources that need normalization, including the Pathway Ontology, which looks to have lots of information. I’ll be working on it via this GitHub issue.
There’s also PathBank, (curated by Yojana but not appearing in the original ComPath publication), BioCyc, MetaCyc, and many others. Each must first be included in PyOBO as I described in a previous post before getting into curation, so I can reuse all the code.
I’m not super happy with any of the names I’ve given to relationships in this
post, either. I’m open to suggestion for improvement. We alternatively discussed
using skos:broader and skos:narrower, as well. Further, I’d love to see
these kinds of relationships appear in the Relation Ontology itself, but
unfortunately I have not been super successful in petitioning them for
improvements in the past, so I may start another open ontology project focused
on relationships themselves.
This is all part of a greater effort, Biomappings, which Ben and I have been working on to make it much easier to curate equivalences and related mappings. I’ll have more to say about that in a future post.